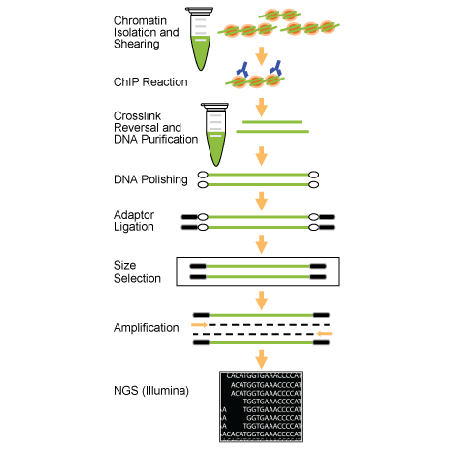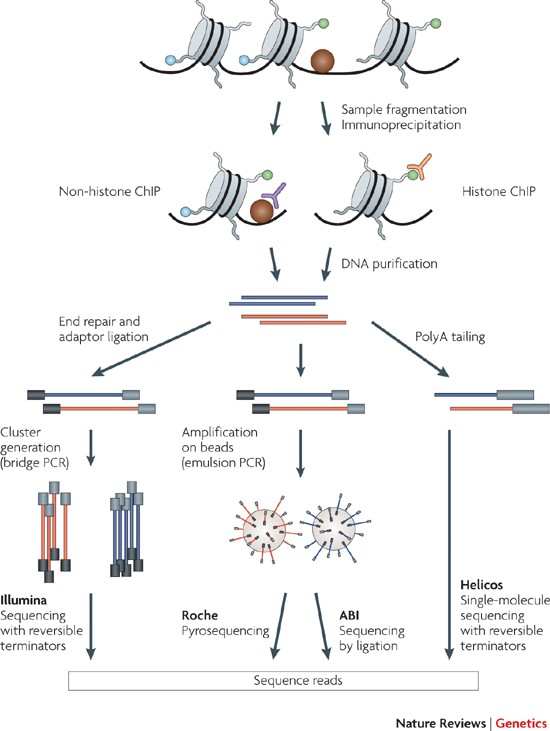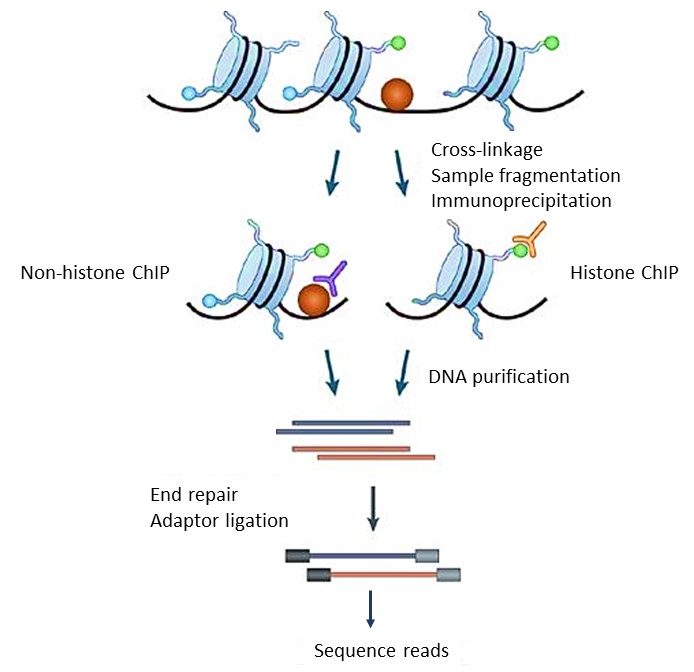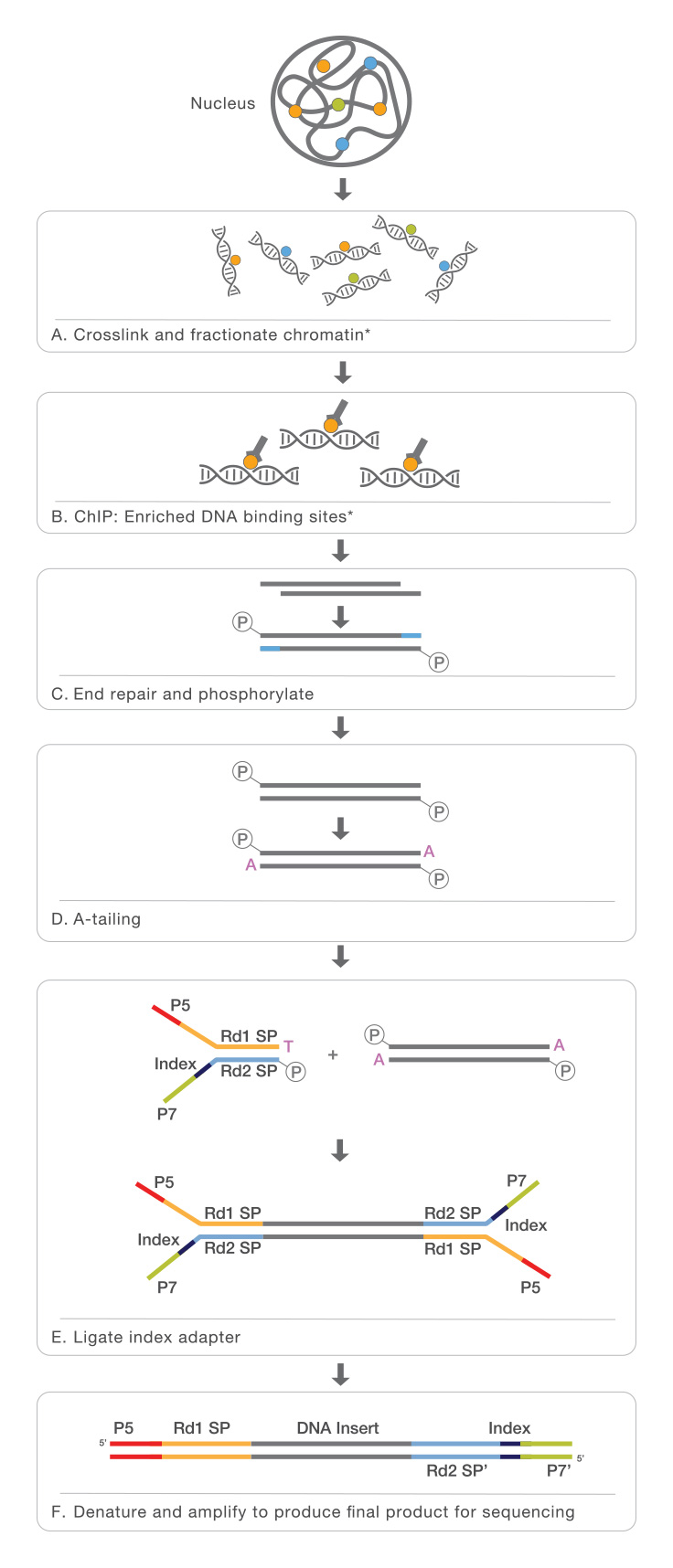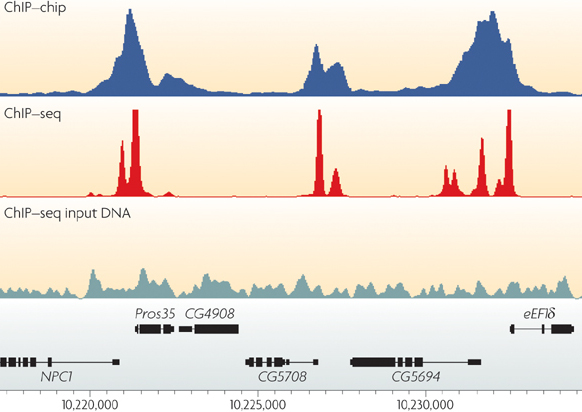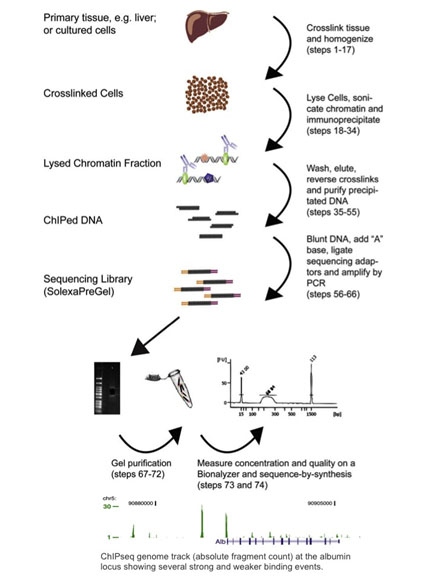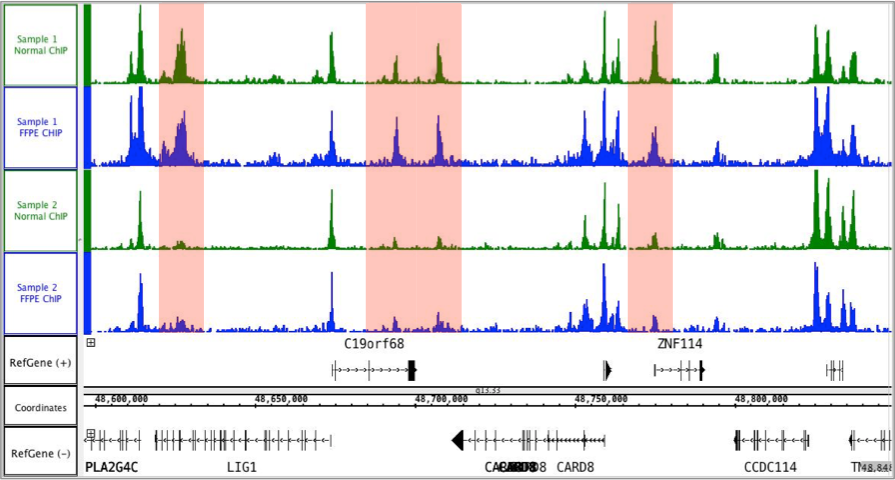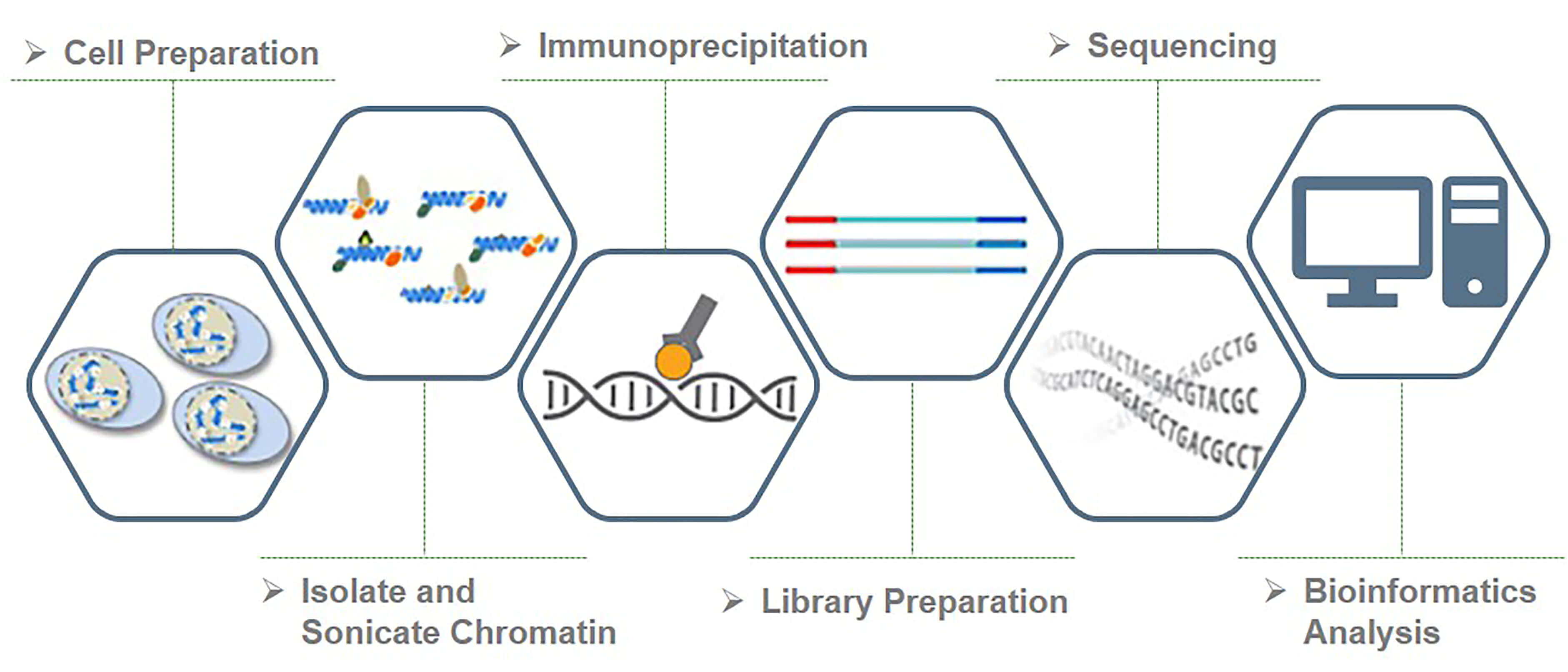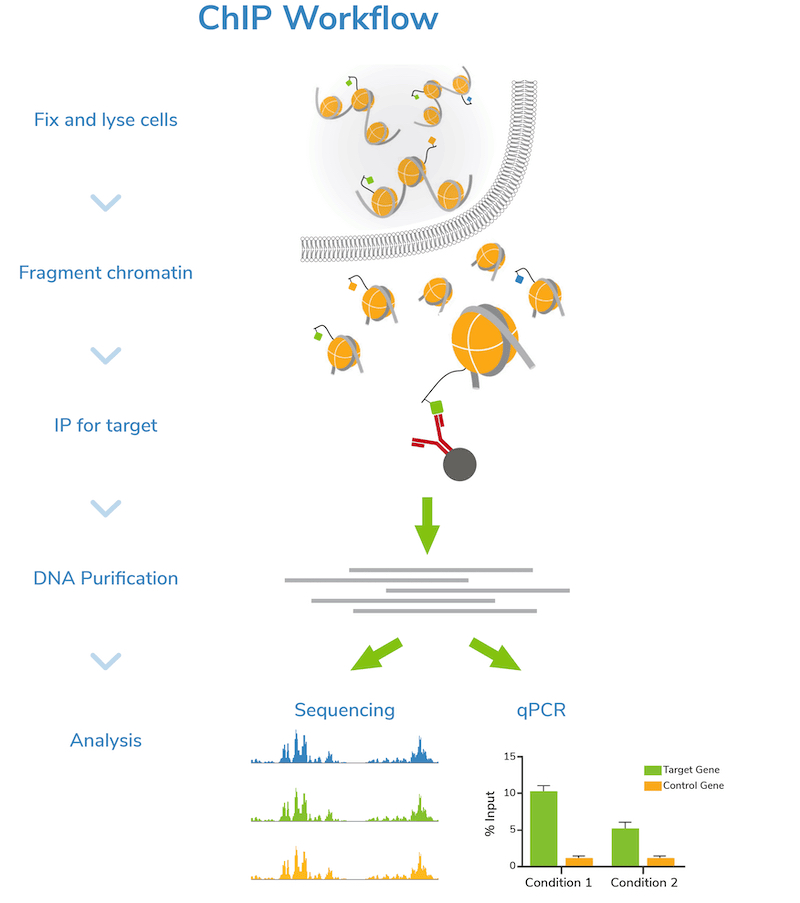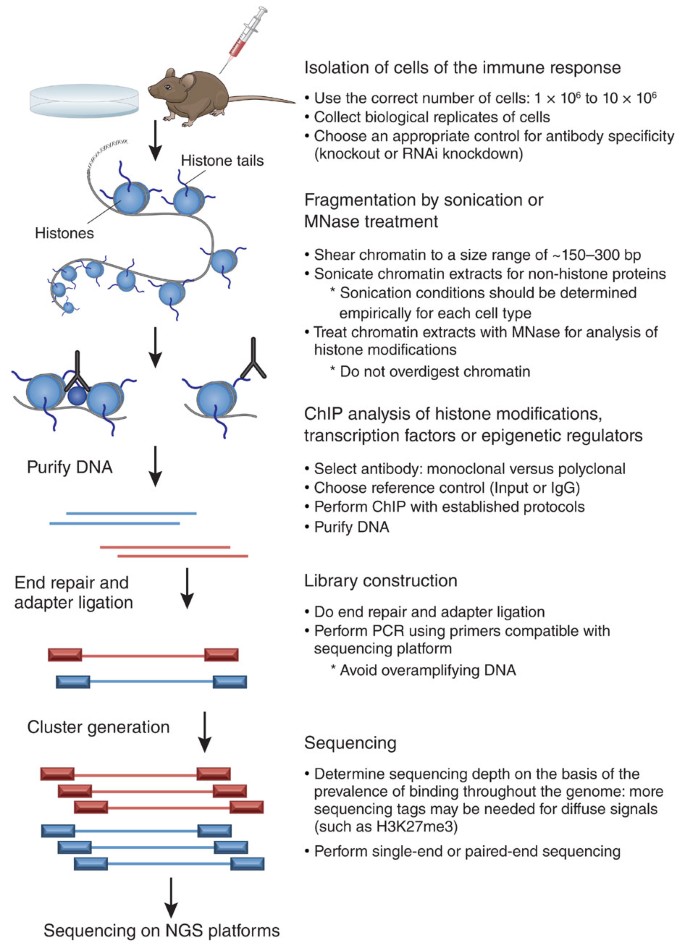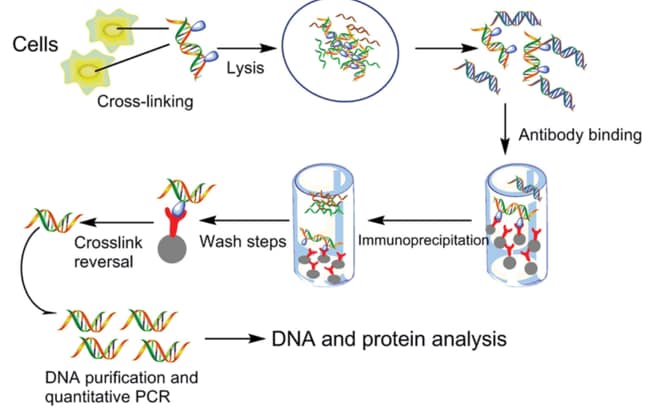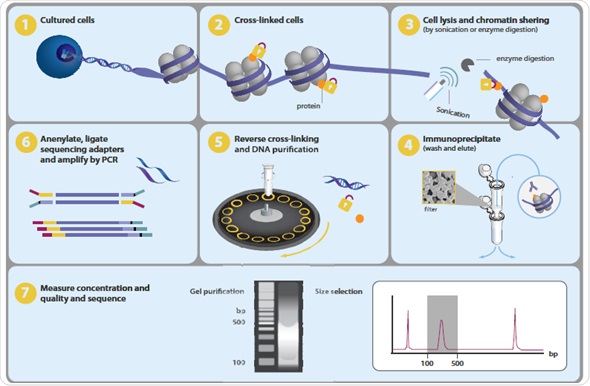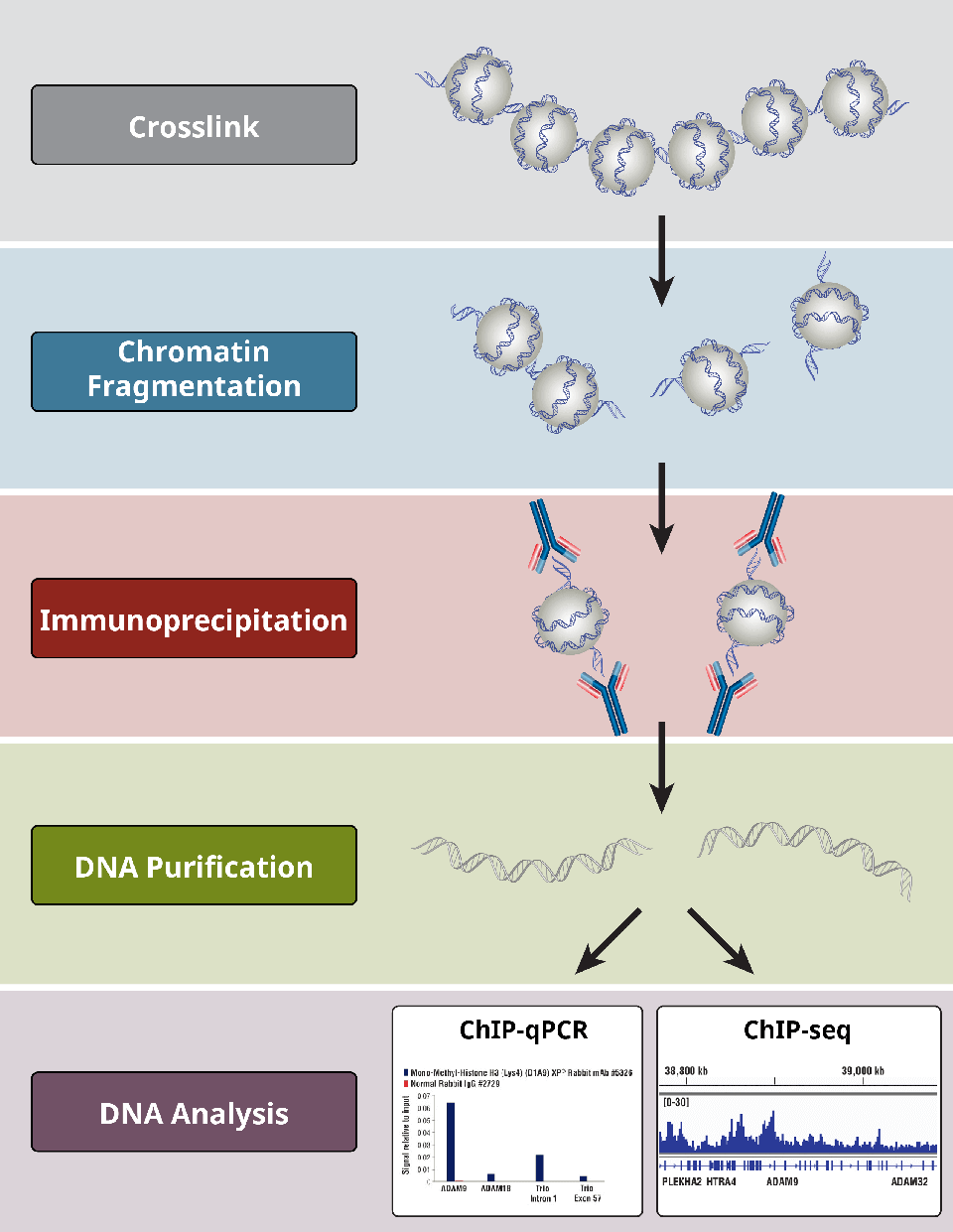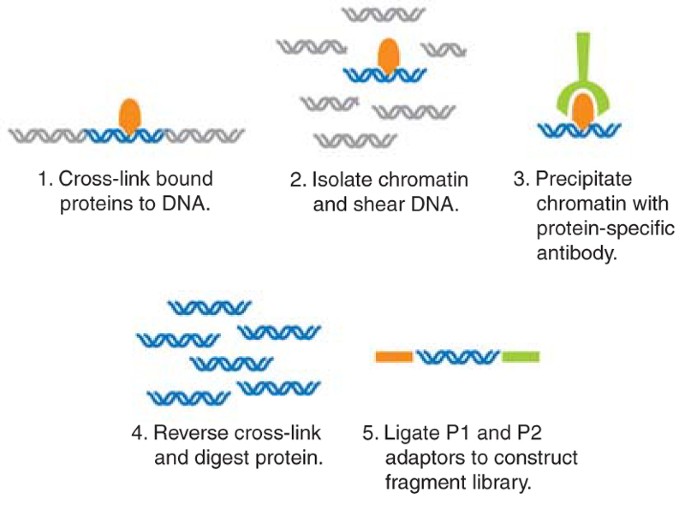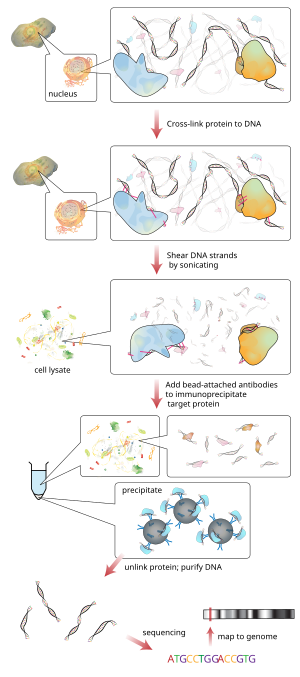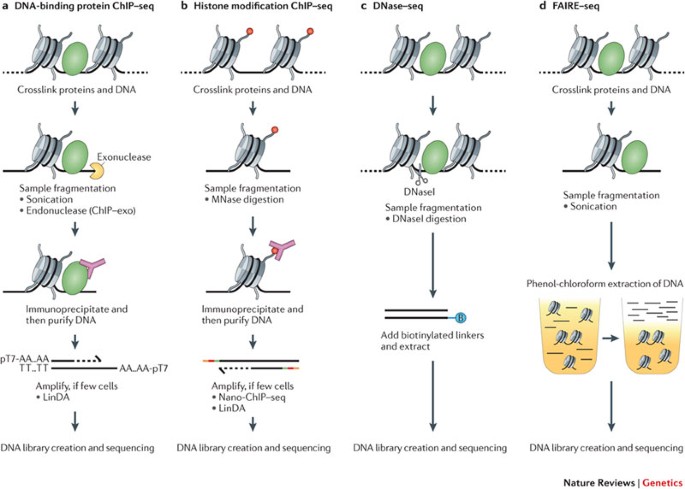
ChIP–seq and beyond: new and improved methodologies to detect and characterize protein–DNA interactions | Nature Reviews Genetics
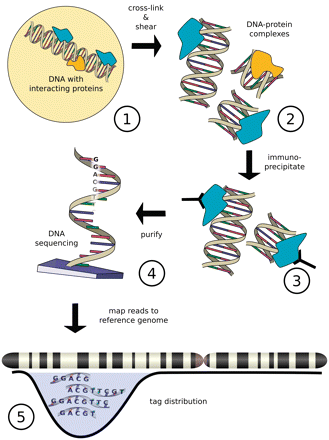
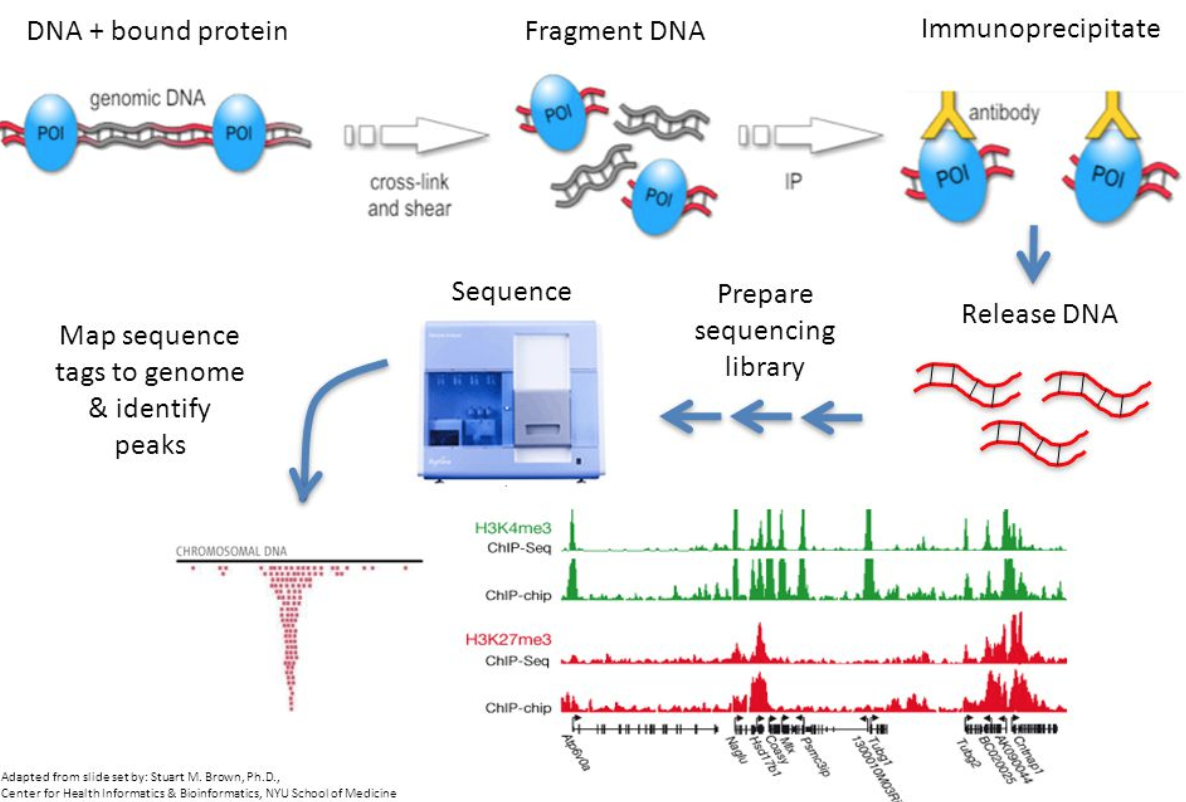
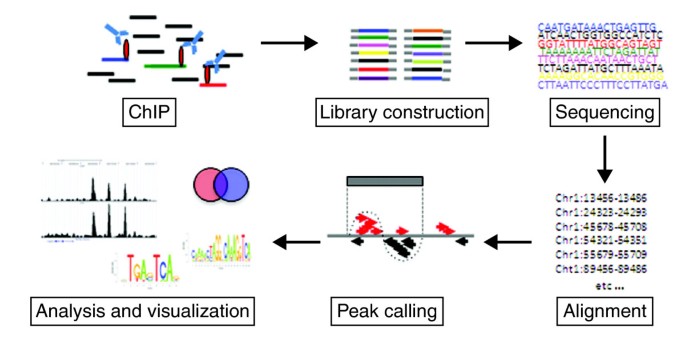
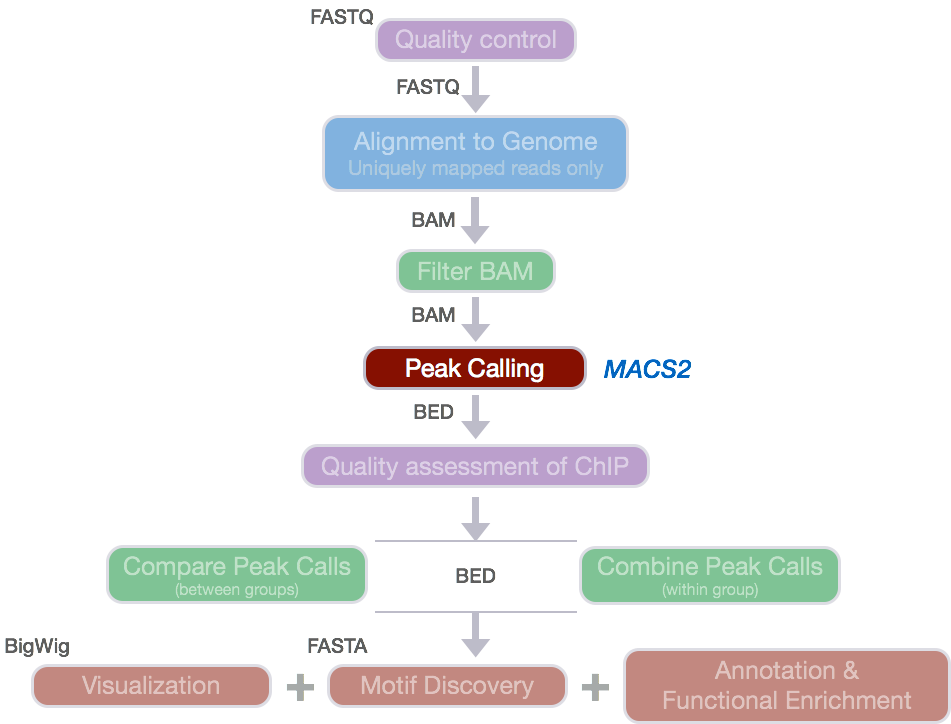
Introduction to ChIP-seq and directory setup | Introduction to ChIP-Seq using high-performance computing

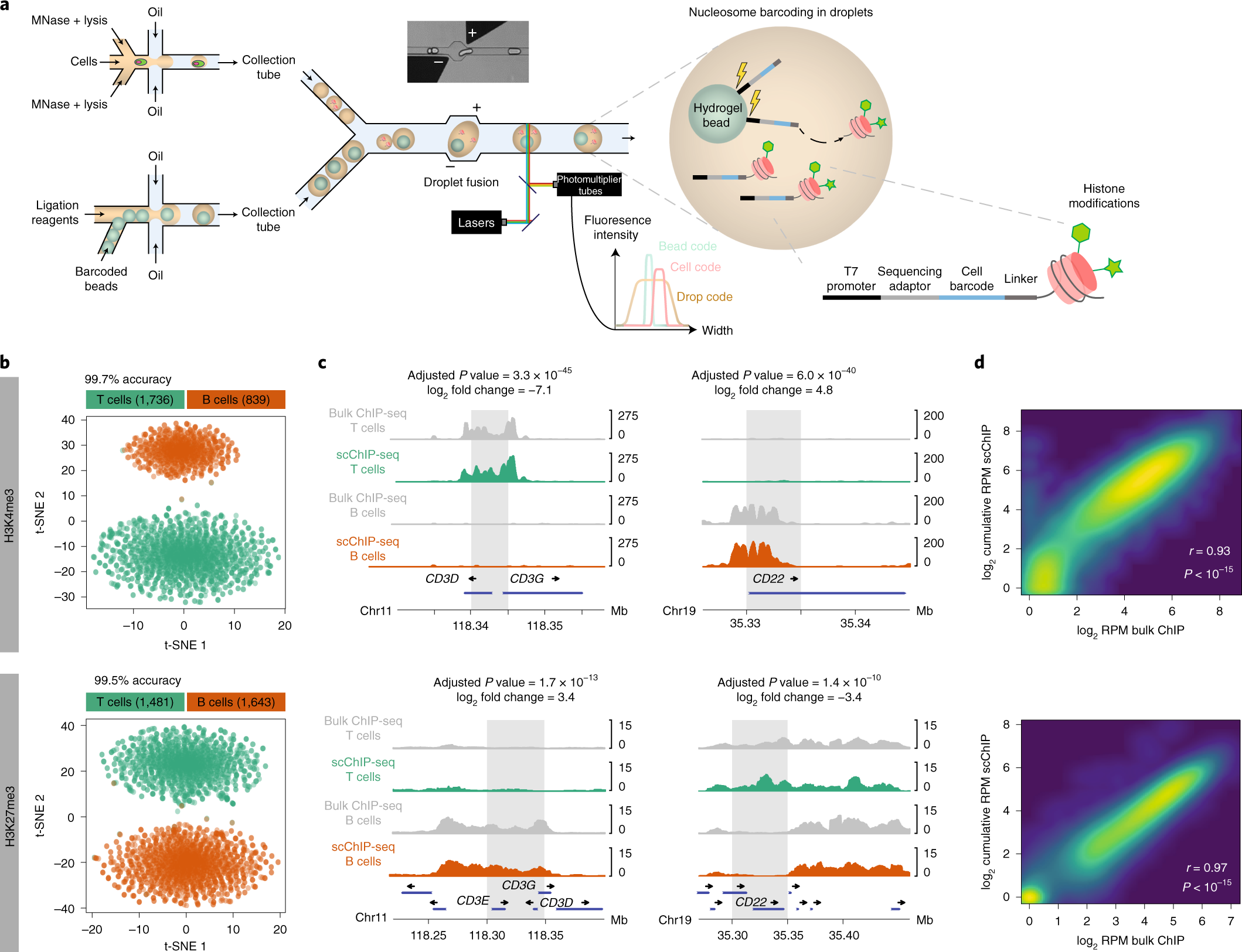
ChIP-seq of plasma cell-free nucleosomes identifies gene expression programs of the cells of origin | Nature Biotechnology