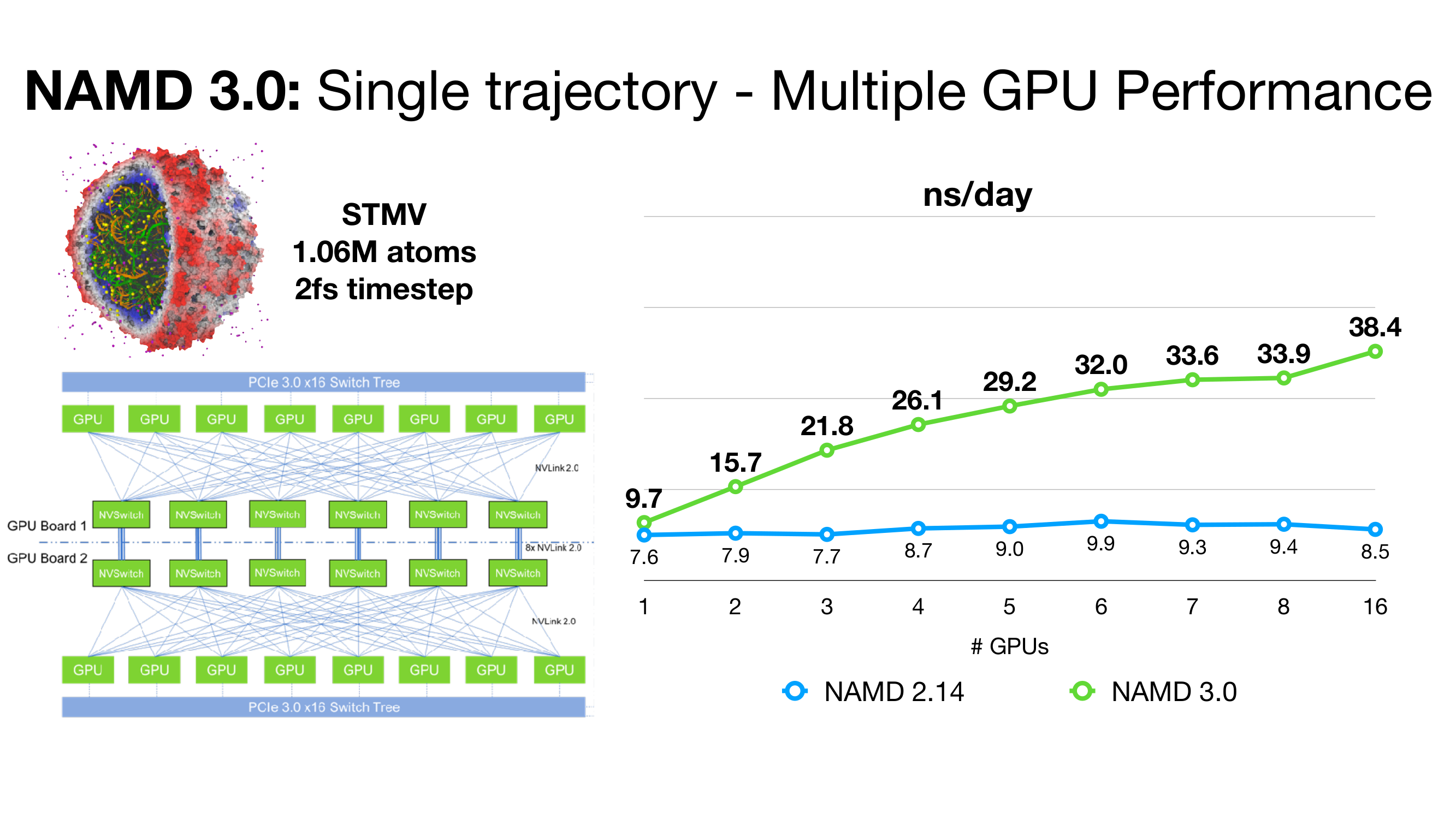
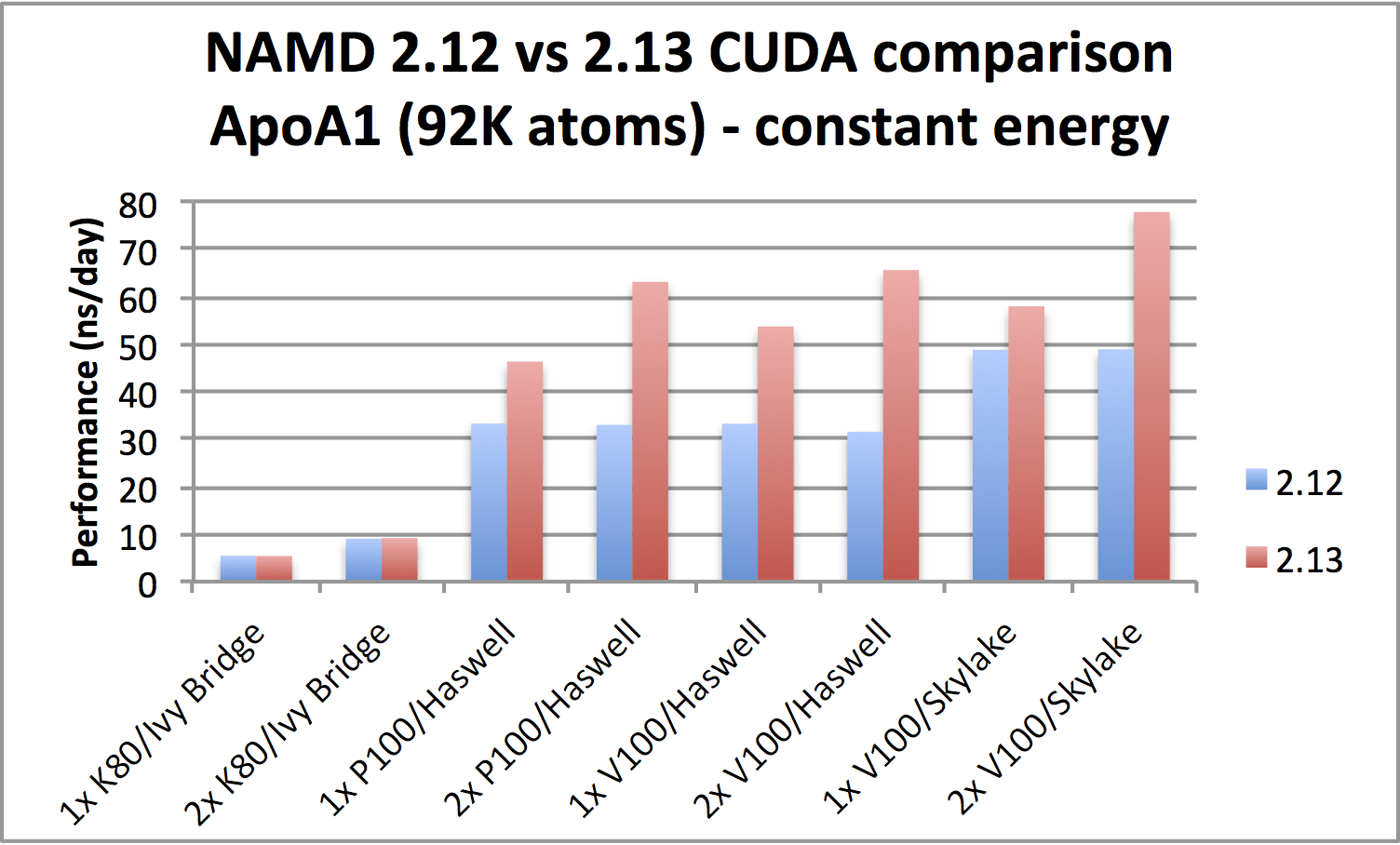
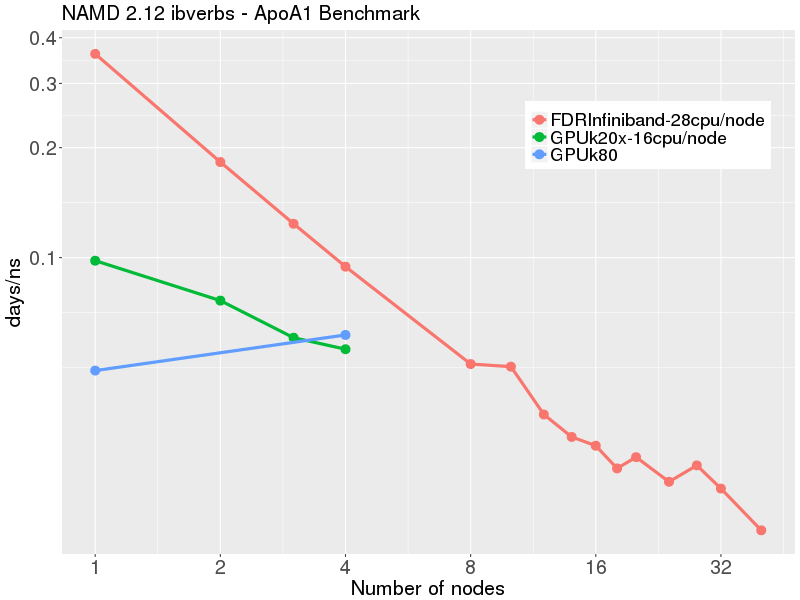
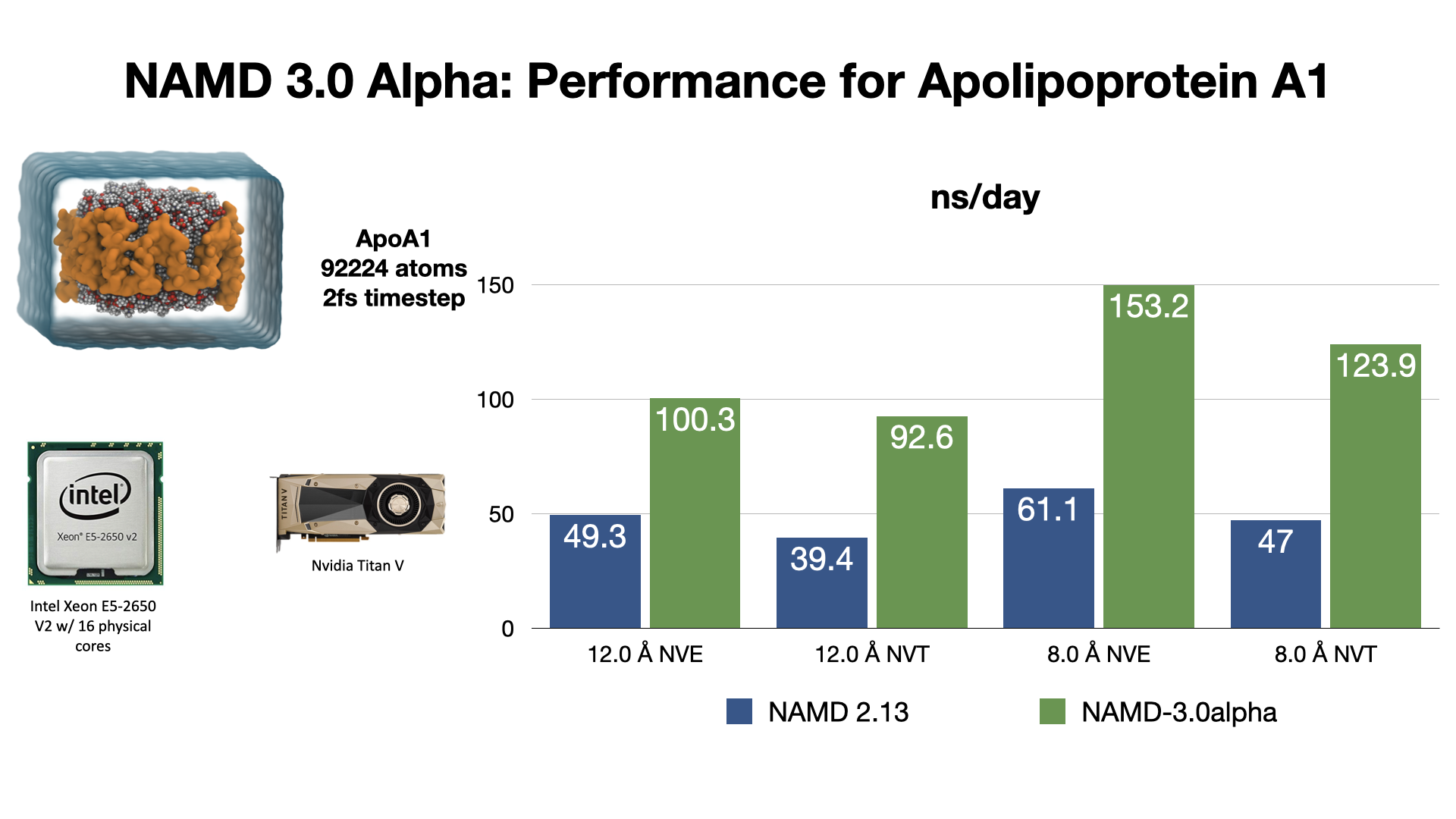
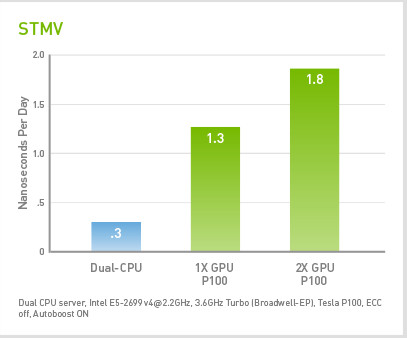
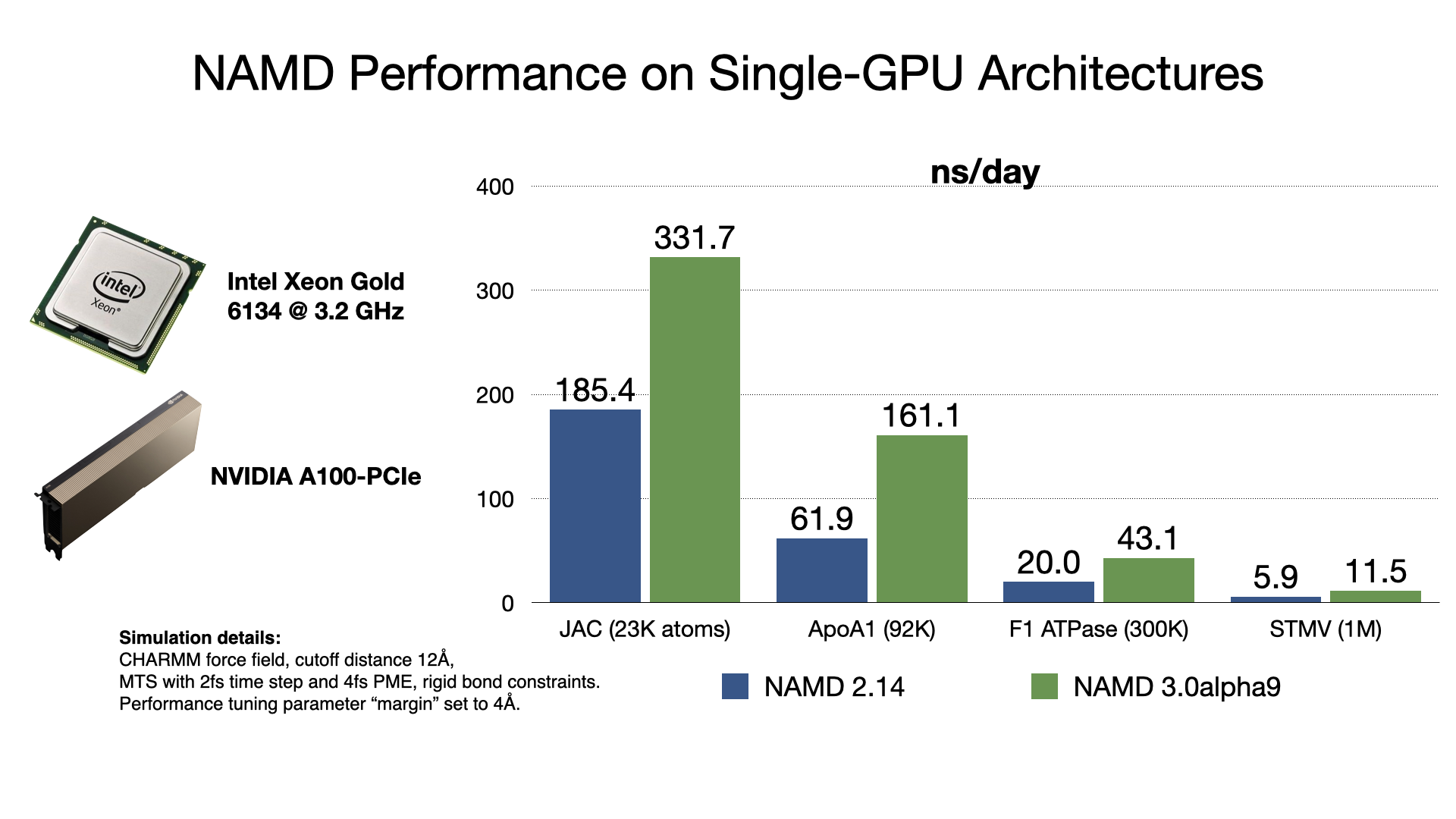
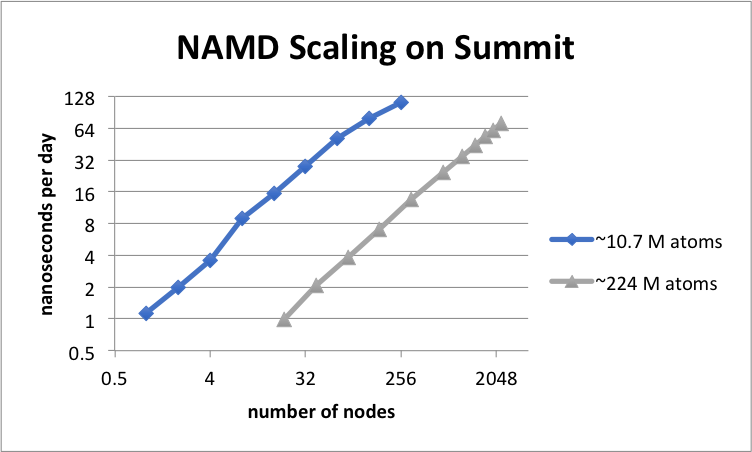
Scalable molecular dynamics on CPU and GPU architectures with NAMD: The Journal of Chemical Physics: Vol 153, No 4

Boosting Free-Energy Perturbation Calculations with GPU-Accelerated NAMD | Journal of Chemical Information and Modeling
A Comparative Performance Ranking of the Molecular Dynamics Software – Running Molecular Dynamics with Amber on Compute Canada