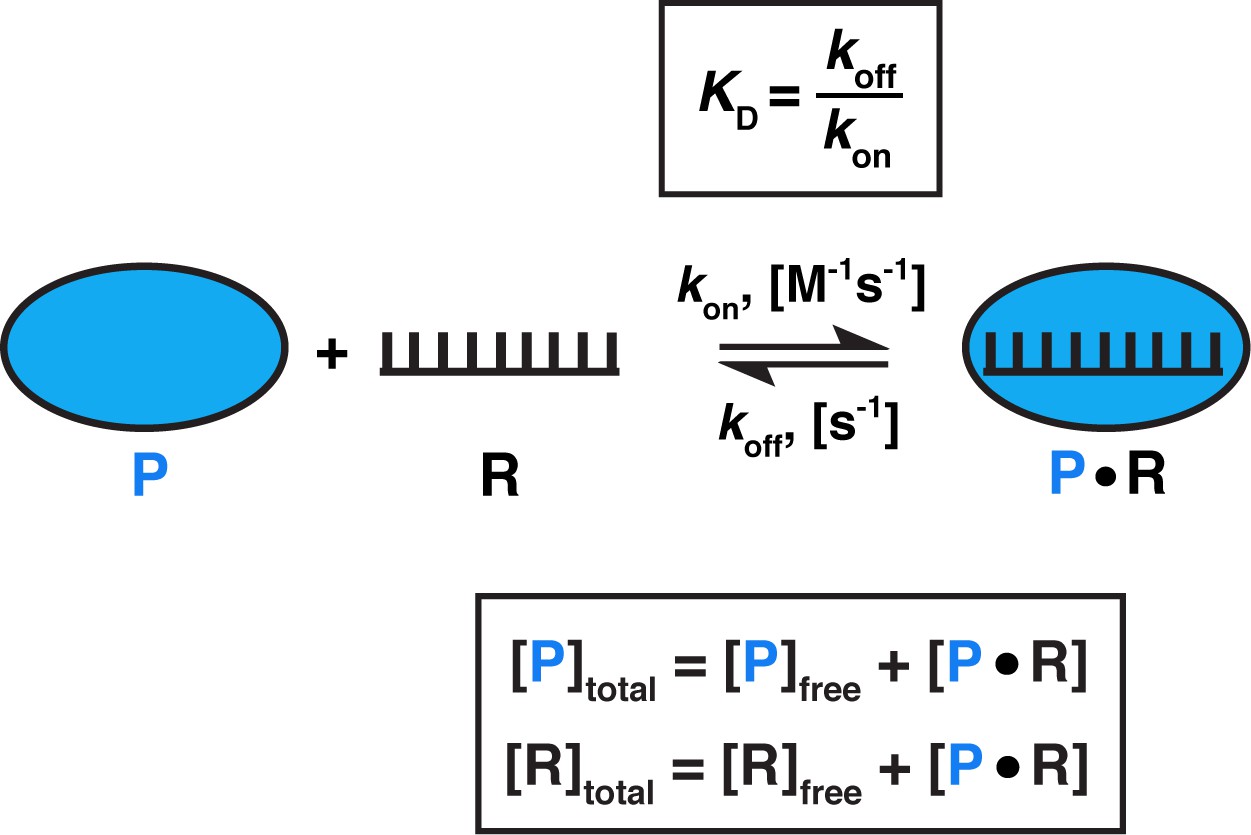
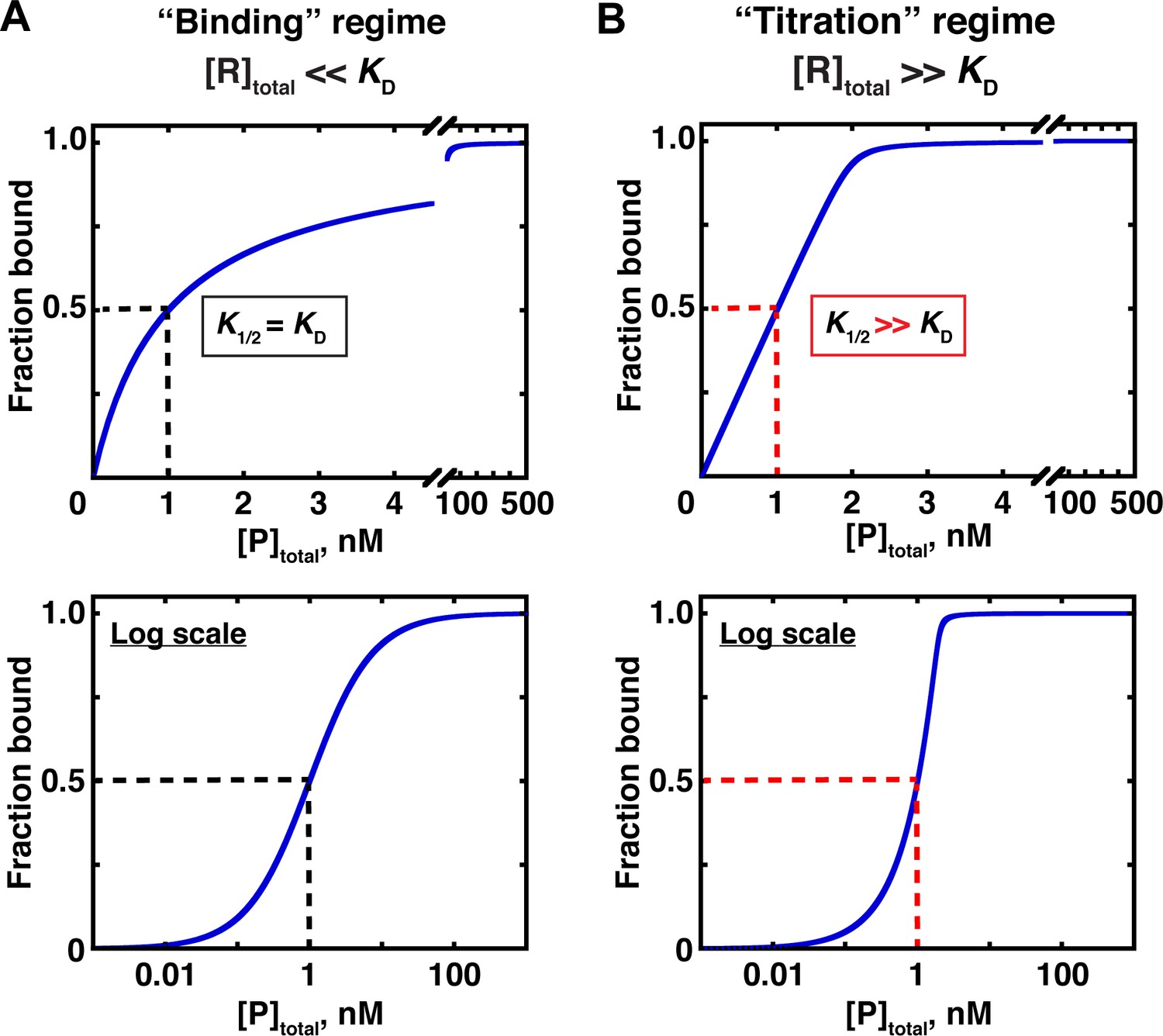
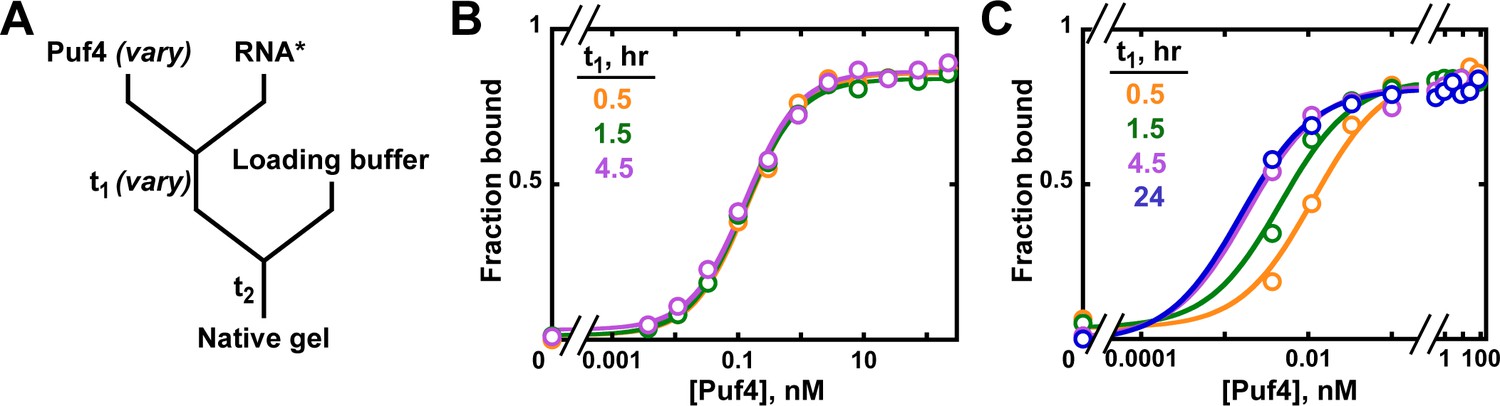
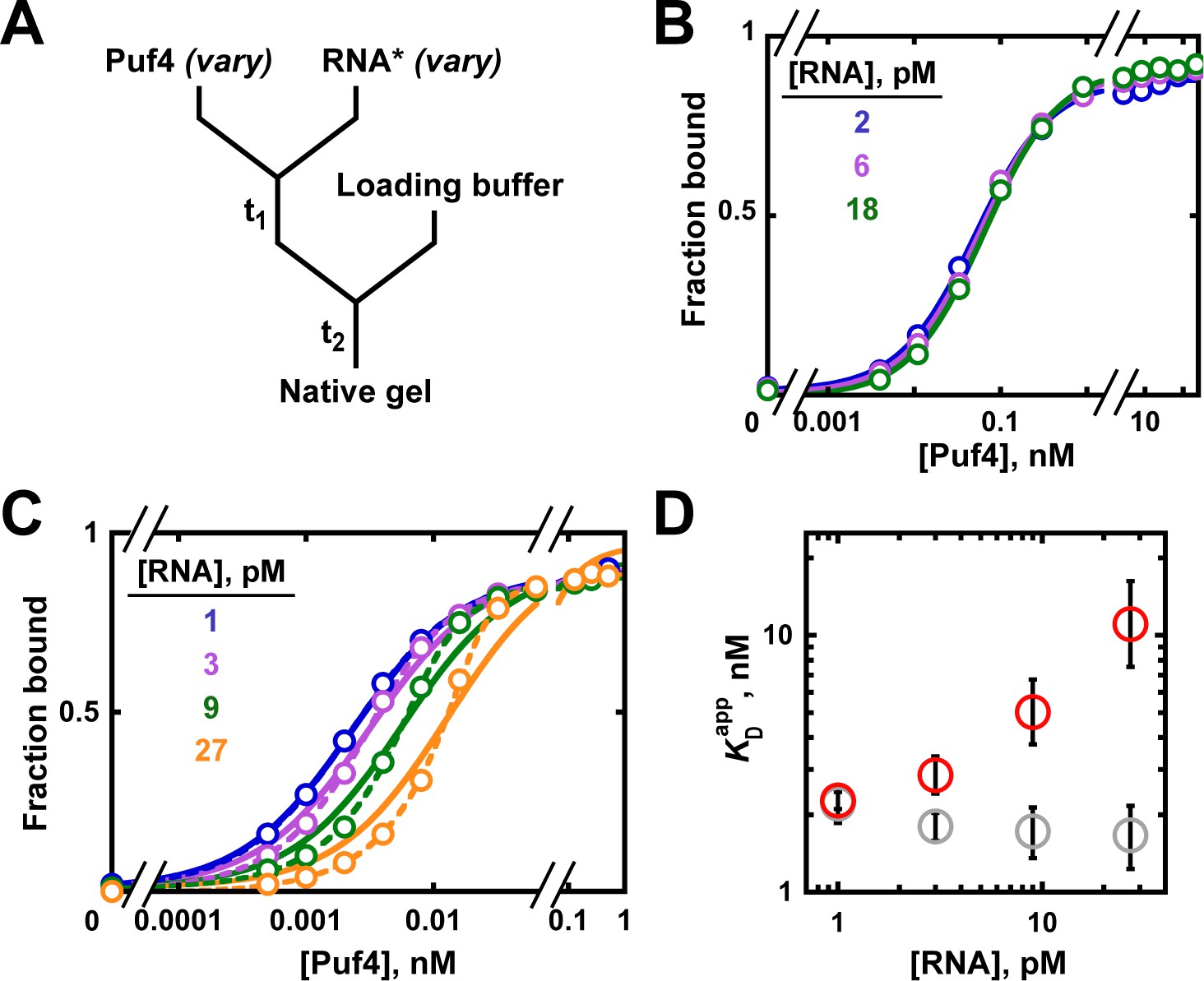
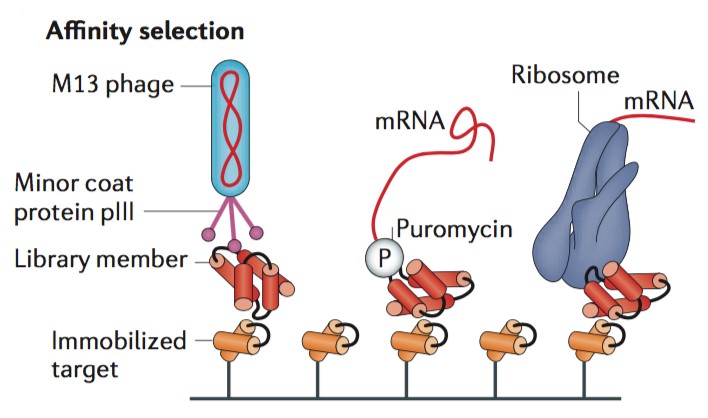
Determining Force Dependence of Two-Dimensional Receptor-Ligand Binding Affinity by Centrifugation: Biophysical Journal
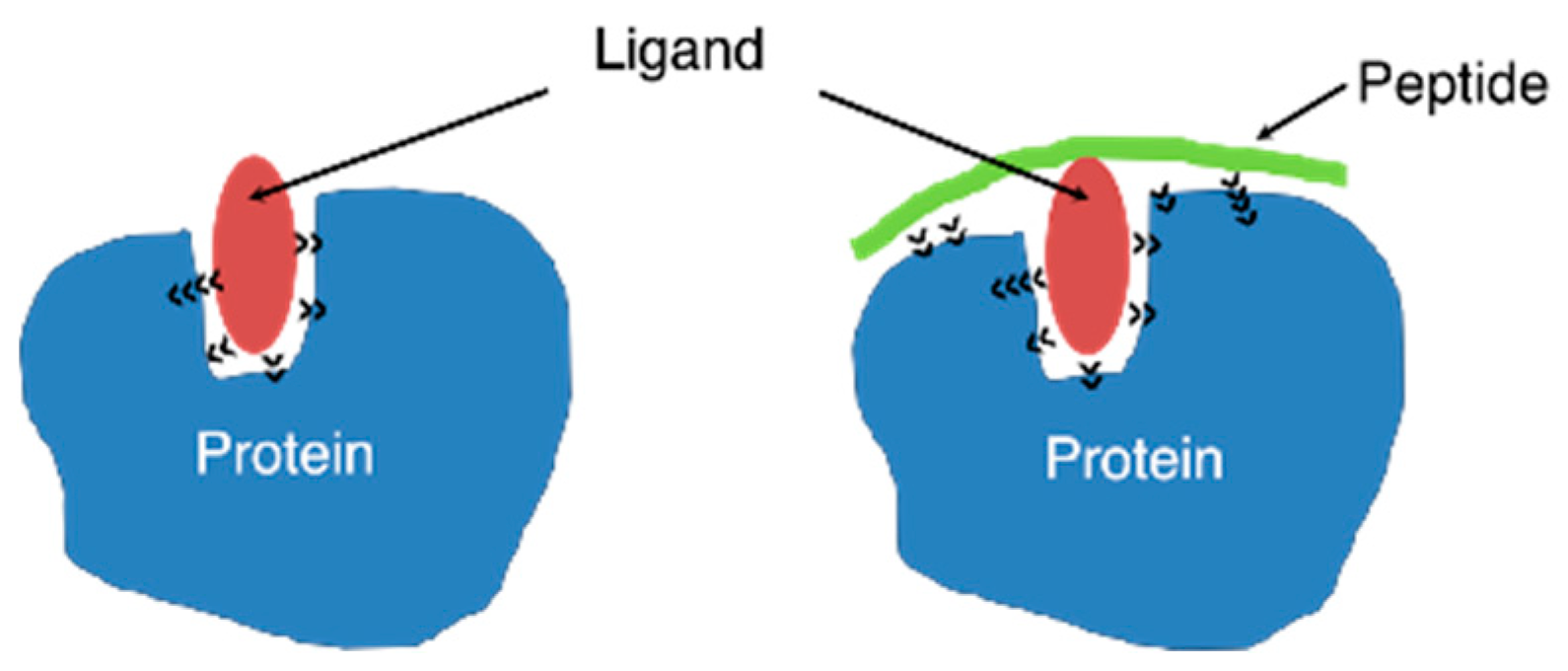
IJMS | Free Full-Text | Enhancement of Binding Affinity of Folate to Its Receptor by Peptide Conjugation

Protein-ligand binding affinity prediction based on profiles of intermolecular contacts - ScienceDirect
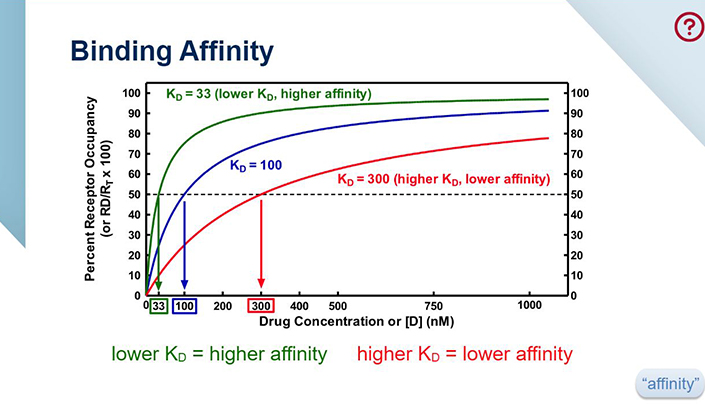
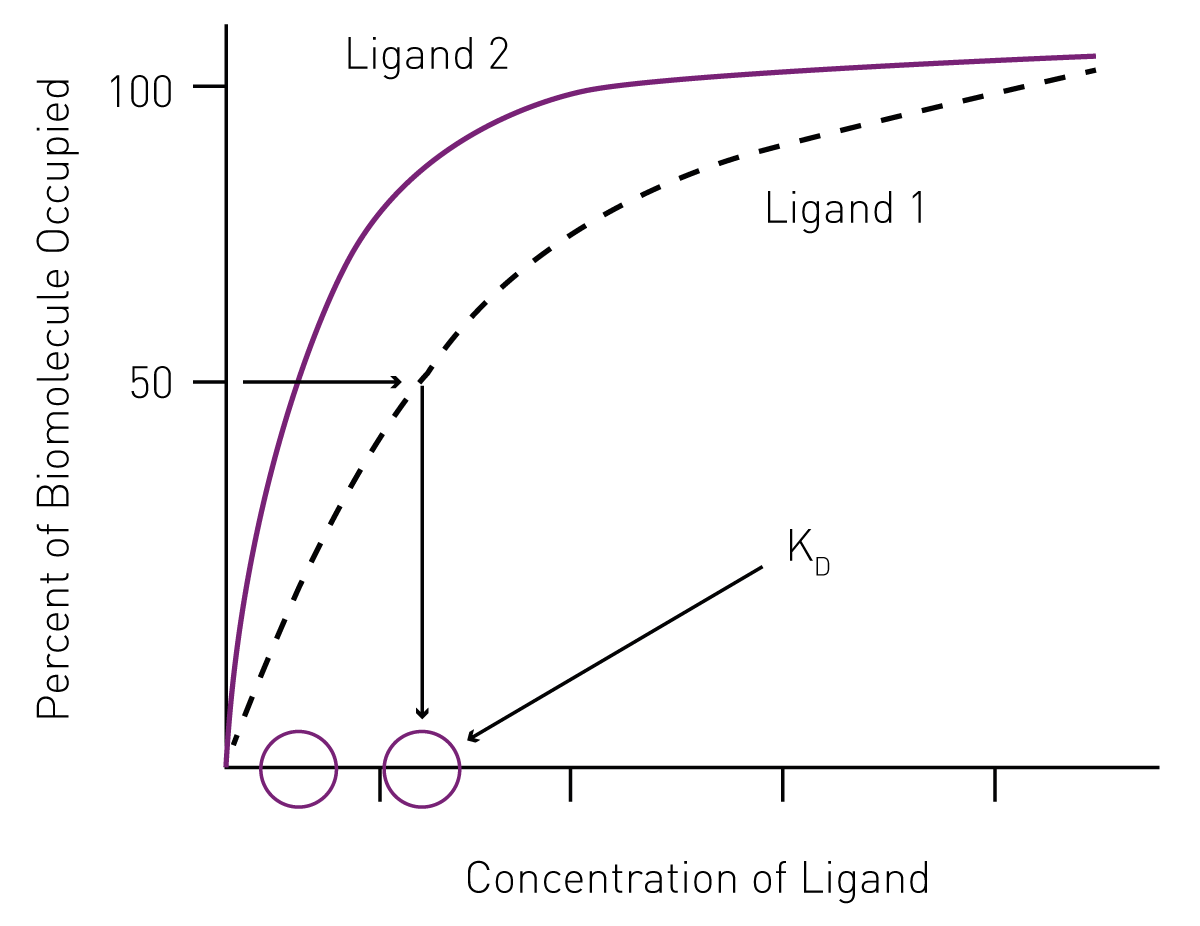
What determines a ligand's affinity for a receptor? Are there multiple factors in play that would make one ligand have a higher affinity than another for a given receptor? - Quora
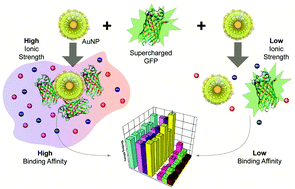
High affinity protein surface binding through co-engineering of nanoparticles and proteins - Nanoscale (RSC Publishing)

Improved Protein–Ligand Binding Affinity Prediction with Structure-Based Deep Fusion Inference | Journal of Chemical Information and Modeling

Binding affinity – “slot/dot blot” filter binding assay for measuring protein/nucleic acid interactions – The Bumbling Biochemist
1: Thermodynamic cycles for calculation of binding affinities using... | Download Scientific Diagram