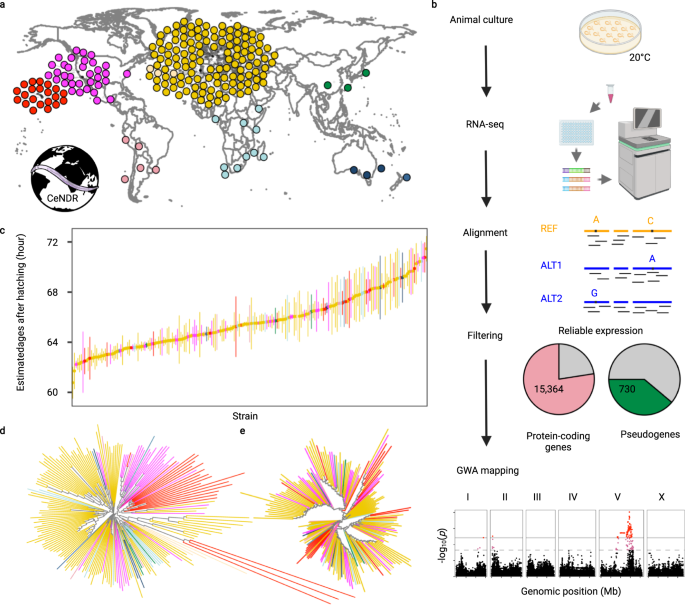
The impact of species-wide gene expression variation on Caenorhabditis elegans complex traits | Nature Communications
The Caenorhabditis elegans homolog of the Evi1 proto-oncogene, egl-43, coordinates G1 cell cycle arrest with pro-invasive gene expression during anchor cell invasion | PLOS Genetics

The multipotency‐to‐commitment transition in Caenorhabditis elegans—implications for reprogramming from cells to organs - Spickard - 2018 - FEBS Letters - Wiley Online Library
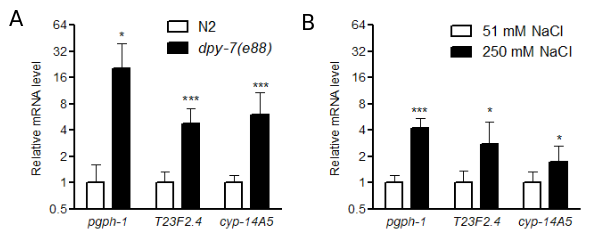
Increased expression of pgph-1, T23F2.4, and cyp-14A5 in C. elegans dpy-7 mutants and by high salt | microPublication

Differential expression of genes in C. elegans reveals transcriptional responses to indirect-acting xenobiotic compounds and insensitivity to 2,3,7,8-tetrachlorodibenzodioxin - ScienceDirect
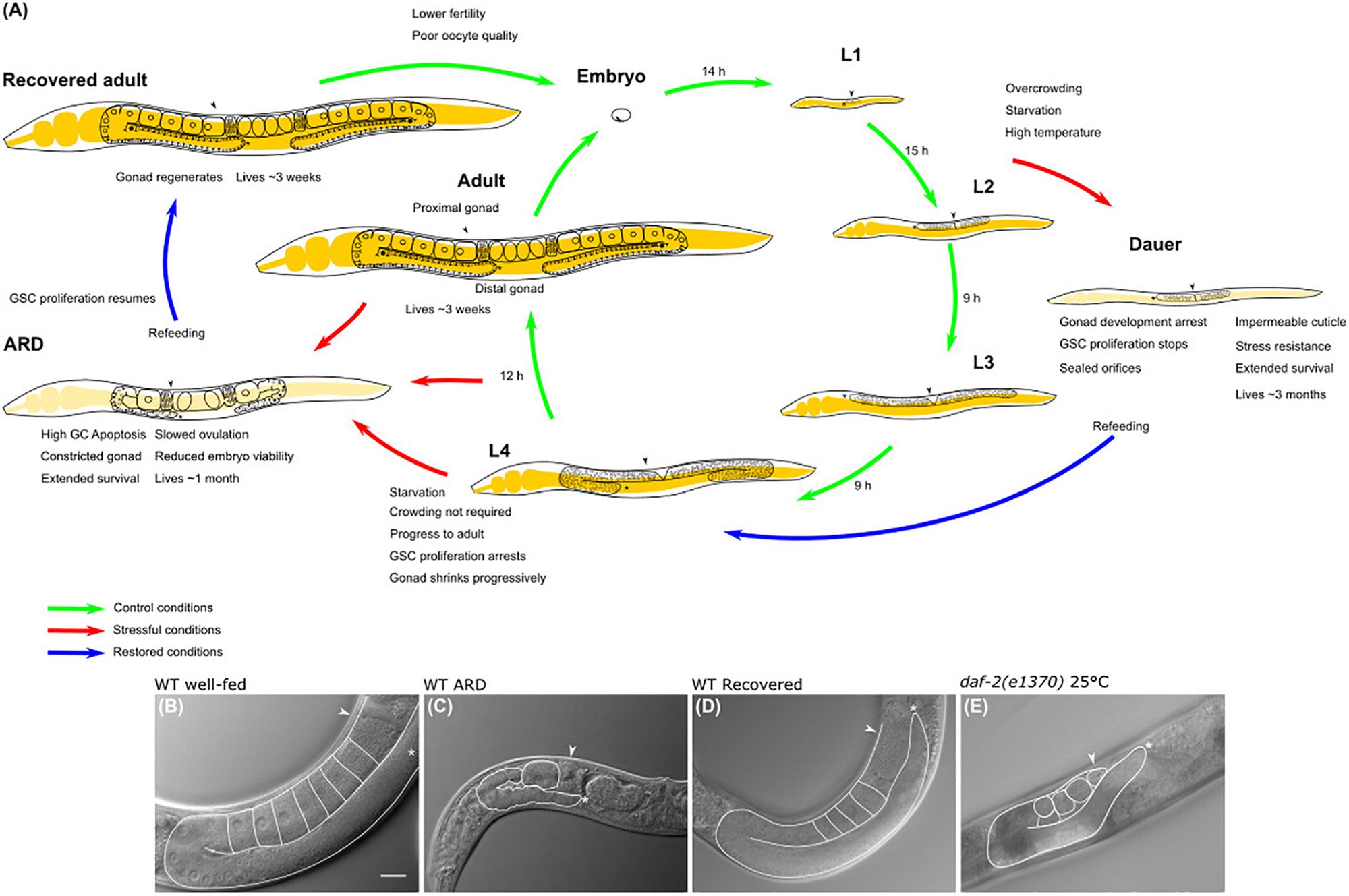
Frontiers | Insights Into the Hypometabolic Stage Caused by Prolonged Starvation in L4-Adult Caenorhabditis elegans Hermaphrodites

Multilevel regulation of muscle-specific transcription factor hlh-1 during Caenorhabditis elegans embryogenesis | SpringerLink

Caenorhabditis elegans saposin-like spp-9 is involved in specific innate immune responses | Genes & Immunity

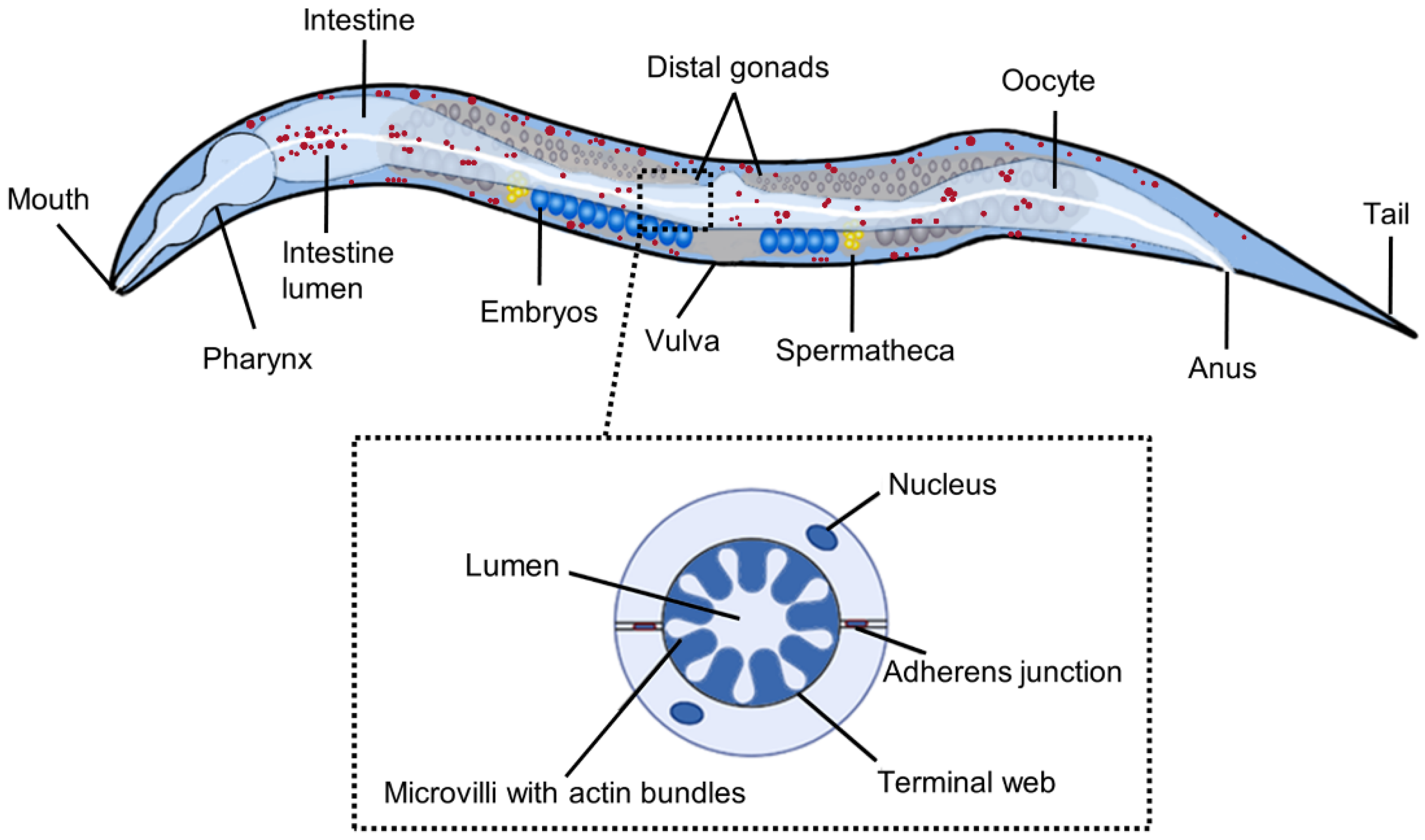
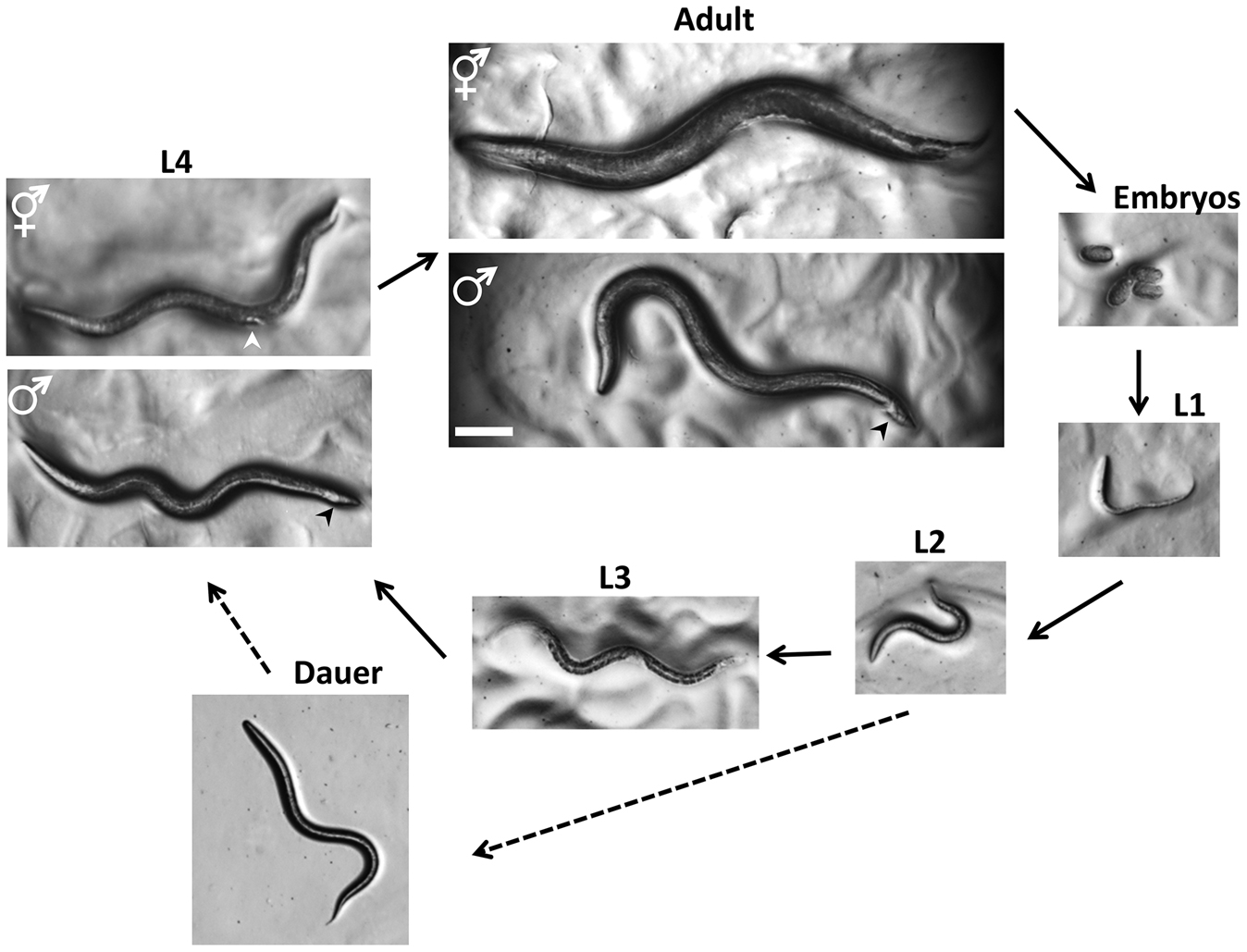
Working with Worms: Caenorhabditis elegans as a Model Organism - Meneely - 2019 - Current Protocols Essential Laboratory Techniques - Wiley Online Library

ERK phosphorylates chromosomal axis component HORMA domain protein HTP-1 to regulate oocyte numbers | Science Advances
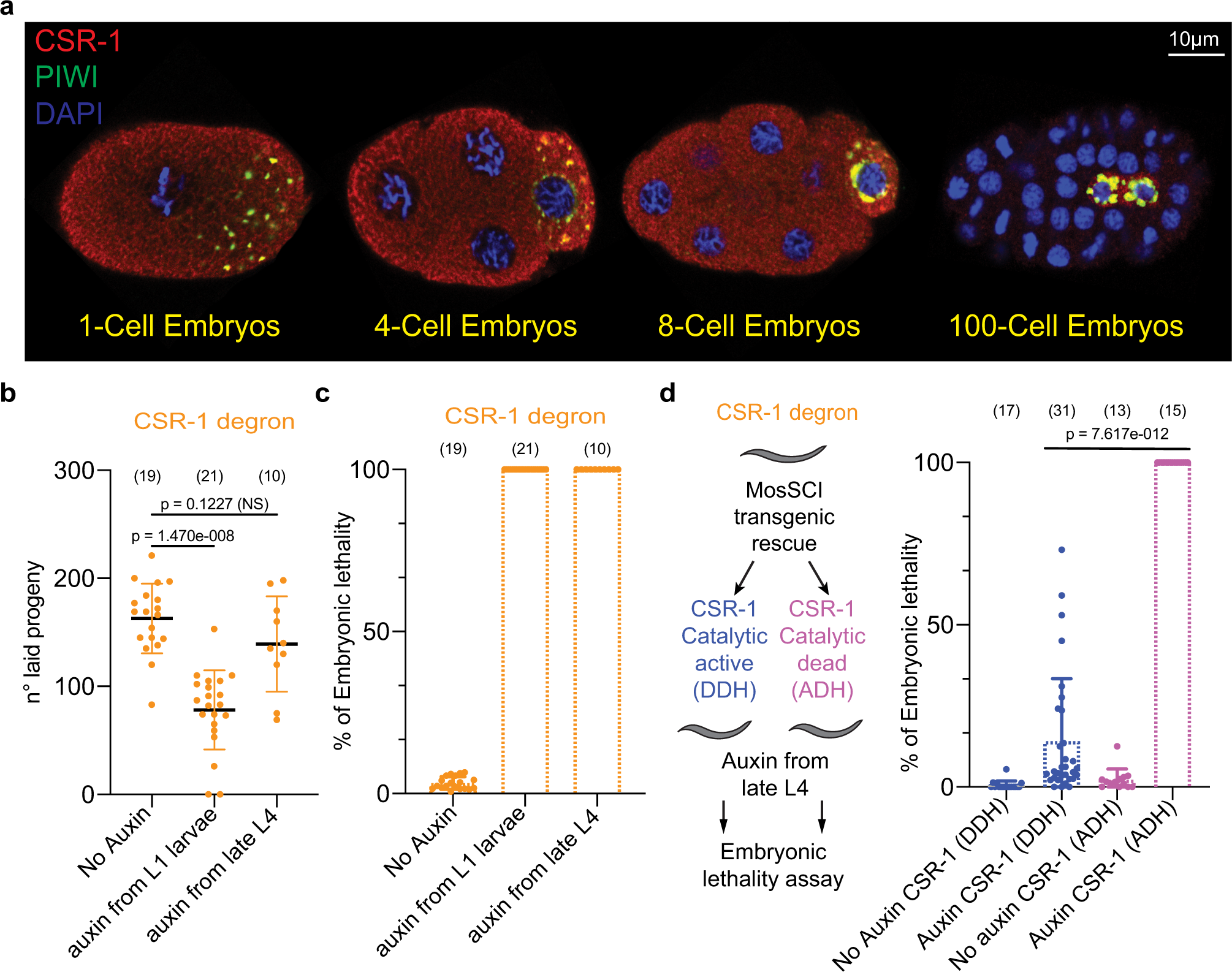
Germline inherited small RNAs facilitate the clearance of untranslated maternal mRNAs in C. elegans embryos | Nature Communications
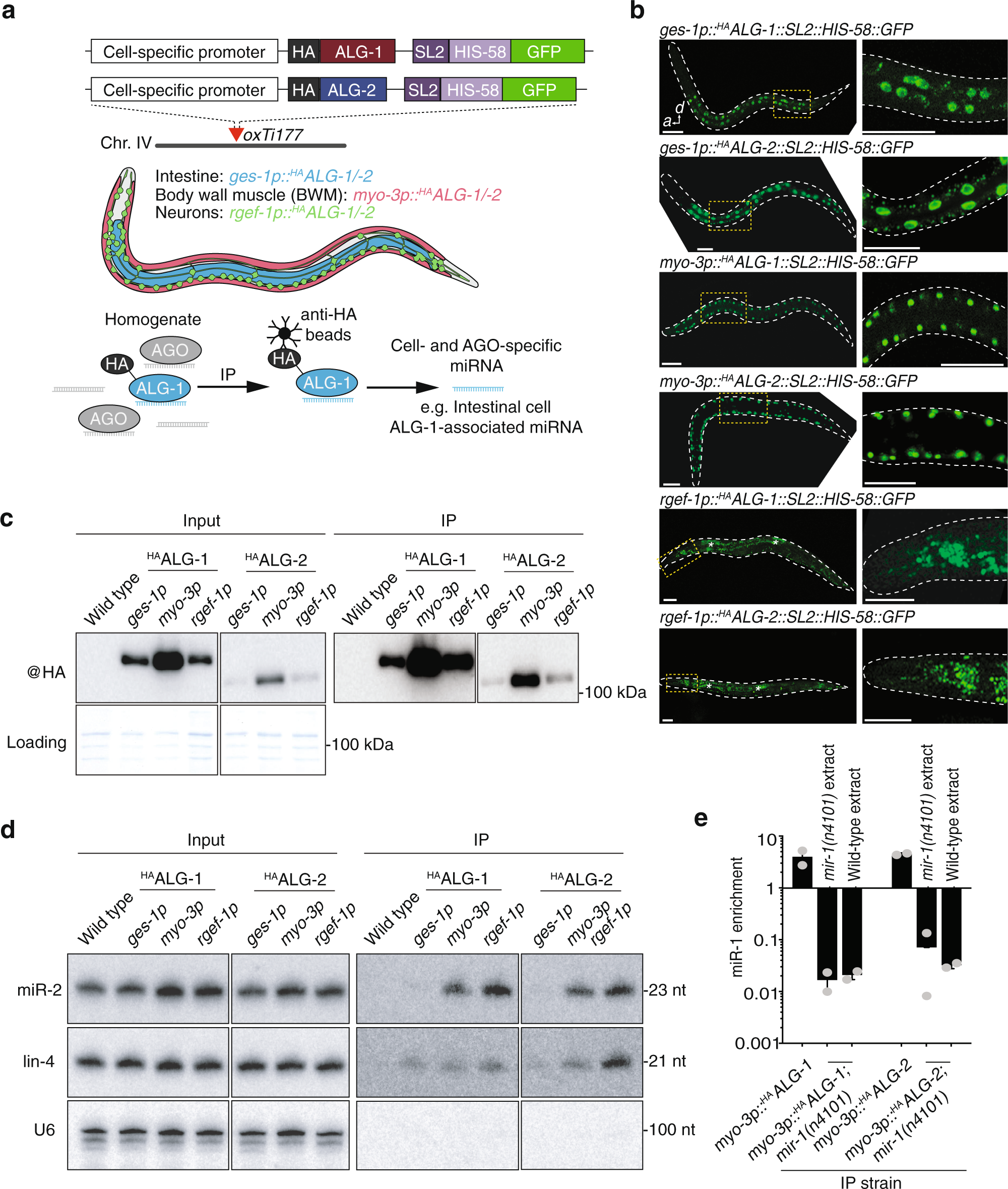
Cell-type-specific profiling of loaded miRNAs from Caenorhabditis elegans reveals spatial and temporal flexibility in Argonaute loading | Nature Communications

In vivo single-molecule imaging identifies altered dynamics of calcium channels in dystrophin-mutant C. elegans | Nature Communications
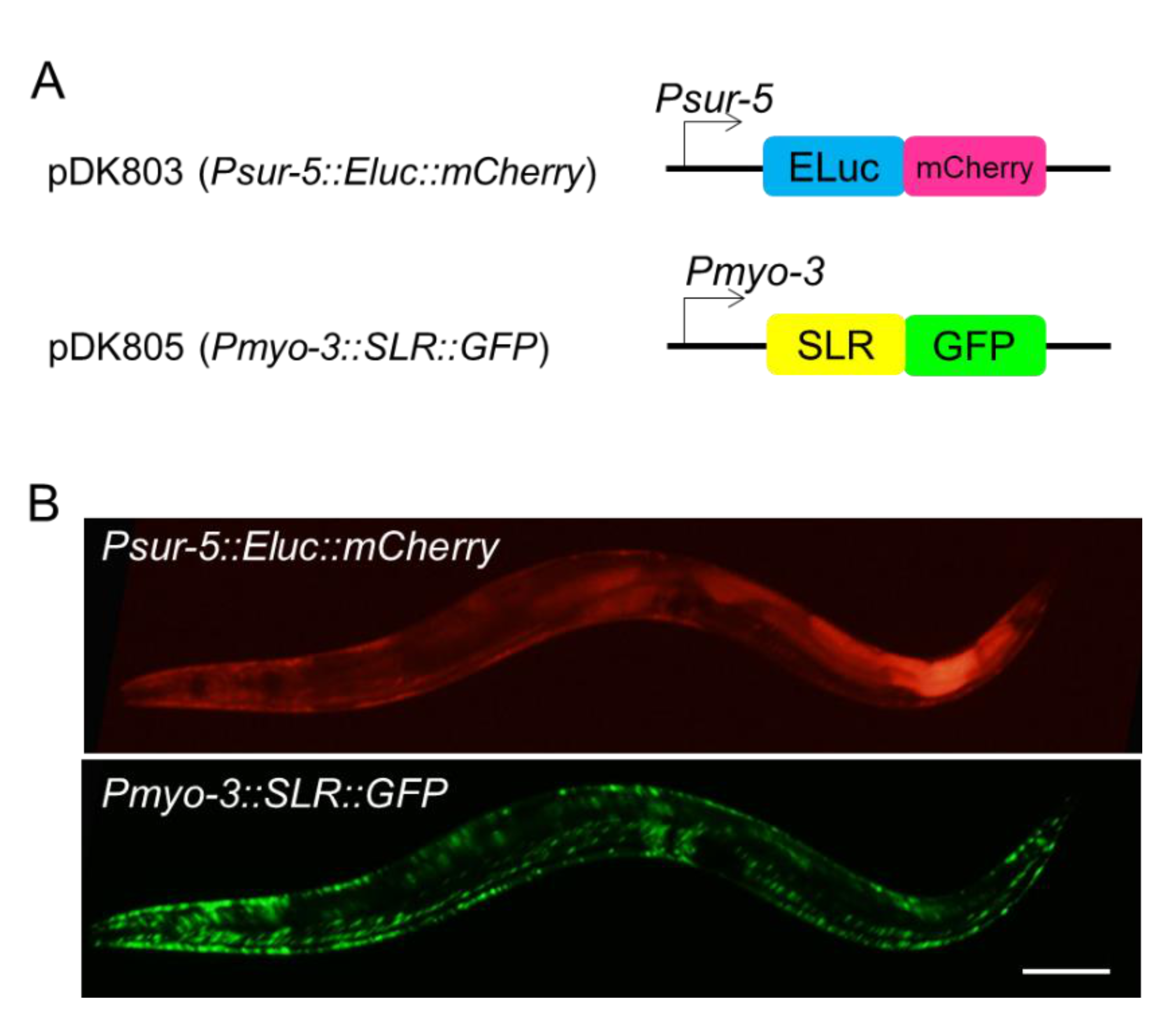
IJMS | Free Full-Text | In Vivo Simultaneous Analysis of Gene Expression by Dual-Color Luciferases in Caenorhabditis elegans